Research
I am an evolutionary geneticist who is interested in general properties of genetic systems more so than the history or function of specific genes.
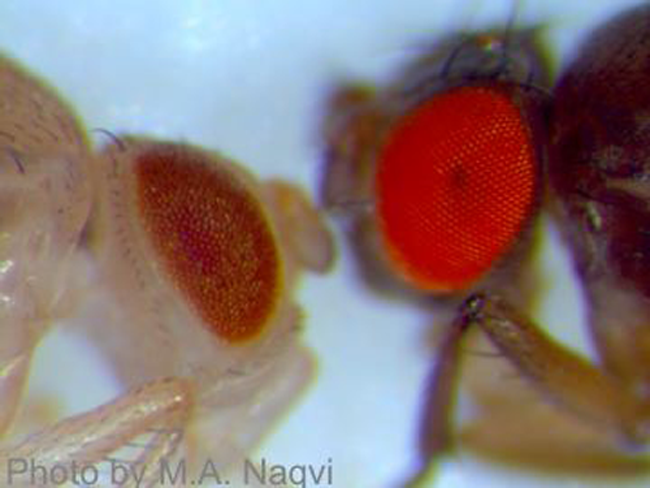

I ask questions such as: Are sexually-derived offspring more fit than asexually-derived offspring (and why)? Do mutation and recombination rates change with condition? If so, why would it matter? To what extent are populations burdened by deleterious alleles? How do ecological and genetic factors affect this load? I approach these problems both theoretically and empirically using several biological systems, including the mighty fruit fly, Drosophila melanogaster (a few cool mutants are pictured here). Our empirical work spans classic experimental approaches to modern evolutionary genomics.
Sex & Genetic Mixing using theory, experiments, and genomics
In theory:Most plants and animals reproduce sexually rather than asexually, but why is this so? Why should organisms continually mix up their genomes? This is one of the great outstanding problems in evolutionary biology. When organisms reproduce sexually, two genomic processes, segregation and recombination, allow for the shuffling of genes and the production of offspring genotypes different from the parental ones. How do these different processes contribute to the evolution of genetic mixing? [PDF] I am interested in understanding the group-level and gene-level advantages of segregation and recombination. Because a variety of different ideas have been put forth as possible solutions to the perplexing problem of “why sex?”, this work has led to an interest in several related areas including mutation load, host-parasite coevolution, and plasticity in genomic processes. Theoretical models allow us to more rigorously evaluate ideas about what maintains sex and what are the consequences of having little or no sex. Theoretical modeling is the first of several approaches we use to study sex.
Most plants and animals reproduce sexually rather than asexually, but why is this so? Why should organisms continually mix up their genomes? This is one of the great outstanding problems in evolutionary biology. When organisms reproduce sexually, two genomic processes, segregation and recombination, allow for the shuffling of genes and the production of offspring genotypes different from the parental ones. How do these different processes contribute to the evolution of genetic mixing? [PDF] I am interested in understanding the group-level and gene-level advantages of segregation and recombination. Because a variety of different ideas have been put forth as possible solutions to the perplexing problem of “why sex?”, this work has led to an interest in several related areas including mutation load, host-parasite coevolution, and plasticity in genomic processes. Theoretical models allow us to more rigorously evaluate ideas about what maintains sex and what are the consequences of having little or no sex. Theoretical modeling is the first of several approaches we use to study sex.
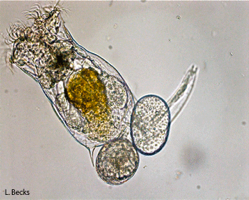
With rotifers: Lab studies allow us to test elements of the theory in real systems held under well-controlled (though artificial) conditions. Several years ago we began an experimental program using the facultatively sexual rotifer Brachionus calyciflorus(pictured here) to test theoretical predictions about the types of conditions that favour increased rates of sex. For example, we have recently experimentally confirmed a classic theoretical prediction that higher rates of sex are favoured during adaptation [PDF]. Using additional manipulations that dissect the population genetic mechanisms favouring sex. These results revealed that sex is favored, as theory predicts, because high-sex alleles become associated with adaptive alleles rather than because sex is creating beneficial gene combinations per se. We are continuing to use rotifers to test other aspects of theory for the maintenance of sex as well as theory for the consequences of evolving without any sex.
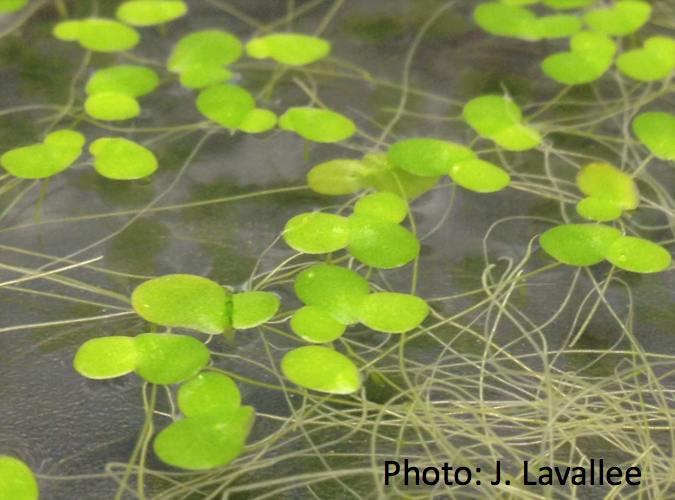

With duckweed: Theory and lab experiments illuminate core principles but ultimately we strive to understand selection on sex and the consequences of high vs. low sex in natural systems. In collaboration with Stephen Wright, we have recently begun studying a group of facultatively sexual plants, duckweeds (fast growing small aquatic plants). We are using population genomics to estimate the frequency of sexual reproduction in different populations and in different species. We are also using population genetics inference to quantify the efficiency of selection across the genome to ask whether populations or species that are more sexual show higher rates of adaptation and/or a lower frequency of deleterious alleles. As part of grounding our population genomics inference, we are doing experiments to measure rates of recombination, mutation, and gene conversion. In addition to genomics, we will be investigating the consequences of sex on fitness in the field. Our aim is to not only infer the population genetic mechanisms maintaining sex but we hope to be able to gain insights into the ecological agents of selection through manipulative field experiments.
Evolutionary Biology of Deleterious Mutations
Mutation is the ultimate source of genetic variation but the vast majority of new mutations are deleterious. These deleterious mutations reduce mean fitness, a phenomenon known as mutation load (see review). Mutation load can be substantial. For example, under the simplest assumptions, classic theory predicts that deleterious mutations will reduce mean fitness by more than 60% if, on average, there is a single deleterious mutation per genome per generation. Because of the potential for mutations to have such large effects on mean fitness, biologists have postulated that deleterious mutations may be a key factor underlying a wide variety of important phenomena including population extinction, the maintenance of genetic variance, the evolution of sexual reproduction, and the evolution of genome size and complexity. Because deleterious alleles affect the health of individuals and reduce the productivity of populations, deleterious alleles are also important with respect to applied issues such as public health, conservation, and the management of biological resources. I am interested in understanding the mutational process itself and whether we can learn general principles about how selection eliminates deleterious alleles.
Mutation rates and spectra
Only recently has it become possible to get good estimates of the mutation rate and we know little about variation in the mutation rate. For both evolutionary and public health reasons, I have been particularly interested in the possibility that individuals in poor health may have elevated rates of mutation. Using fruit flies, we have found that females in low condition have a reduced capacity to repair DNA damage (available from PLoS here) and that low- vs. high-condition flies use different types of repair pathways to fix double strand breaks [PDF]. In another study, we found evidence that flies with low quality genotypes appear to experience elevated rates of mutation [PDF]. We are now using genomic data to discover what types of mutations (e.g., base pair substitutions, indels) occur more often in low- vs. high-quality flies.
Selection on deleterious mutations
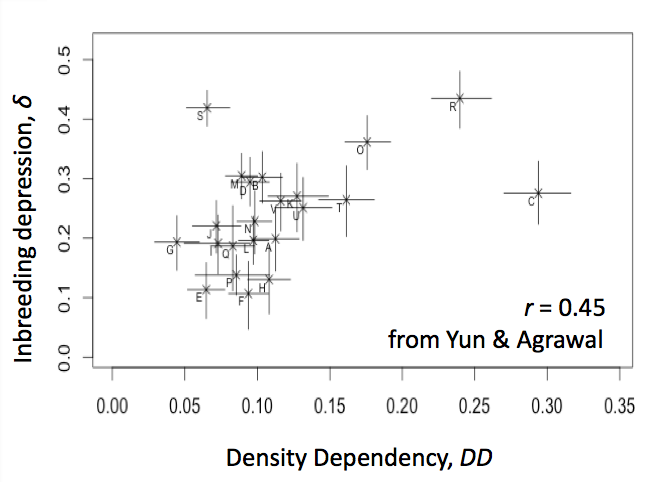
The frequency of deleterious alleles depends on the rate they enter a population by mutation but also on the strength of selection by which they are removed. There are many examples demonstrating that selection on particular mutations depends on context but my group is interested in whether there are there general principles about selection. Are certain environments more selective than others? If so, why? How much do mutational effects depend on the genetic background in which they are measured? Does it matter whether the genetic background is adapted to the environment or not? In a recent study we found that selection against a set of mutations in flies was affected by the environment but not whether the genetic background was adapted to that environment or not [PDF]. From a literature review [PDF], we found little support for the idea that stress increases mean selection but there was indirect evidence that density-dependent competition could be important. Motivated by this, we recently measured selection against inbred genotypes (i.e., inbreeding depression) in 22 environments, characterized both by stress and density dependence, finding the latter was a better predictor of selection (Yun and Agrawal, in press).
Sexual selection and deleterious mutations
Most mutations are likely to be deleterious in both males and females. In males, much of the fitness effect can occur through reduced mating success. For this reason, sexual selection can be a powerful force helping to purge deleterious alleles. Indeed, we have found that selection in flies is stronger through males than females on phenotypic marker mutations [PDF] and spontaneous mutations [PDF]. Furthermore, we have found that male fitness is more sensitive to condition than female fitness across several different assay environments varying in the availability of key resources [PDF]. Results such as these suggest that sexual selection should help eliminate deleterious alleles but others have found evidence sexual selection can hinder natural selection on females if males preferentially harass good quality females. With Howard Rundle (U Ottawa), we have been exploring whether the arena of mating interactions controls the balance between sexual selection’s positive effects on male selection vs. its negative effects on female selection.
Other interests
My other research interests include the maintenance of genetic variance, sexual antagonism and the evolution of sexual dimorphism, host-parasite coevolution, and speciation genetics. In addition to these topics, I am interested in many other aspects of population genetics.